Relations in the Gene Ontology
Overview
The ontologies of GO are structured as a graph, with terms as nodes in the graph and the relations (also known as object properties) between the terms as edges (more ontology information at Gene Ontology Overview). Just as each GO term is defined, the relations between GO terms are also categorized and defined. This document provides a description of some of the commonly used relationships in GO: is a (is a subtype of); part of; has part; regulates, negatively regulates and positively regulates.
All terms (except from the root terms representing each aspect) have an is a sub-class relationship to another term; for example,
GO:1904659:glucose transport is a GO:0015749:monosaccharide transport.
The Gene Ontology employs a number of other relations, including part of, e.g.
GO:0031966:mitochondrial membrane is part of GO:0005740:mitochondrial envelope
and regulates, e.g:
GO:0098689:latency-replication decision regulates GO:0019046:release from viral latency
This set is not exhaustive and includes only a subset of relations used in the GO ontologies, logical definitions and annotations. For more technical information about relations and their properties used in GO and other ontologies see the OBO Relations Ontology (RO).
Using relations to group related annotations
Relations are widely used by GO browsing tools such as AmiGO and QuickGO and in over-representation analysis to group related GO annotations. For example, the annotations on the AmiGO page for kinase activity include annotations made to ‘fucokinase activity’, ‘protein kinase activity’ etc. Note that not all relations can be safely used to group annotations via the GO graph. Please see below for notes and examples illustrating when and why grouping of annotations via relationships can be done safely.
Conventions used to describe relations
There are a number of ways of referring to and representing logical relations. The GO relations documentation uses the following conventions (in keeping with the graph-based terminology):
- A node refers to a GO term
- A parent refers to the node closer to the root(s) of the graph, and a child to that closer to the leaf nodes; for the relations is_a and part_of, the parent would be a broader GO term, and the child would be a more specific term
- The arrowhead indicates the direction of the relationship
- Dotted lines represent an inferred relationship, i.e. one that has not been expressly stated in the ontology.
This diagram would be interpreted as follows:
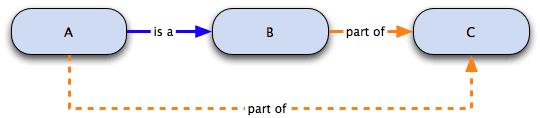
- A is a B
- B is part of C
- we can infer that A is part of C
The formal mathematical / logical representation of the inference made in the graph above is: is a ∘ part of → part of
GO basics
Nodes in the GO graph can have any number and type of relationships to other nodes. Like hierarchies—for example, a family tree or a taxonomy of species—a node may have connections to more than one child (more specific) node, but unlike them, it can also have more than one parent (broader) node, and different relations to its different parents; for example, a node may have a part of relationship to one node, and an is a relationship to another. The following diagram illustrates this point:
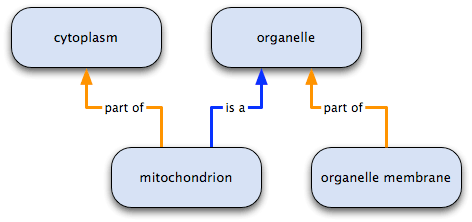
- mitochondrion has two parents: it is an organelle and it is part of the cytoplasm;
Main relations used in GO
| Relation | Description | Use for grouping annotations | |
|---|---|---|---|
| is a | The is a relation forms the basic structure of GO. If we say A is a B, we mean that node A is a subtype of node B. For example, mitotic cell cycle is a cell cycle, or lyase activity is a catalytic activity. | It is safe to use is a to group annotations. For example if a gene product X is annotated as having tyrosine kinase activity and the ontology records that tyrosine kinase activity is a (type of) kinase activity, then we can safely conclude that gene product X has kinase activity. | |
| part of | The part of relation is used to represent part-whole relationships. part of has a specific meaning in GO, and a part of relation would only be added between A and B if B is necessarily part of A: wherever B exists, it is as part of A, and the presence of the B implies the presence of A. However, given the occurrence of A, we cannot say for certain that B exists. | It is safe to use part of to group annotations. For example if a gene product X is annotated as located in the inner mitochondrial membrane and the ontology records a part of relation between inner mitochondrial membrane and mitochondrion, we can safely conclude that X is located in a mitochondrion. | |
| has part | The logical complement to the part of relation is has part, which represents a part-whole relationship from the perspective of the parent. As with part of, the GO relation has part is only used in cases where A always has B as a part, i.e. where A necessarily has part B. If A exists, B will always exist; however, if B exists, we cannot say for certain that A exists. i.e. all A have part B; some B part of A. | It is NOT correct to use has part for grouping annotations. For example, we might assert in the ontology that a receptor tyrosine kinase activity has part ATP hydrolysis activity. However it would not then be correct to group all annotations to kinase activity under ATPase activity. | |
| regulates | A relation that describes case in which one process directly affects the manifestation of another process or quality, i.e. the former regulates the latter. The target of the regulation may be another process, for e.g., regulation of a pathway or an enzymatic reaction, or it may be a quality, such as cell size or pH. Analogously to part of, this relation is used specifically to mean necessarily regulates: if both A and B are present, B always regulates A, but A may not always be regulated by B., i.e. all B regulate A; some A are regulated by B. | Unlike is a and part of, grouping annotations to gene products grouped via regulates changes the relationship between the GO term and the gene product over the is a and part of relations. If gene product X is annotated as involved in a process that regulates glycolysis, it would not be correct to conclude that X participates in glycolysis. Nevertheless, some tools use regulates relations to group annotations. This can be useful for gene-set enrichment. The resulting gene sets include genes that are involved in processes that are causally related to the grouping term. |